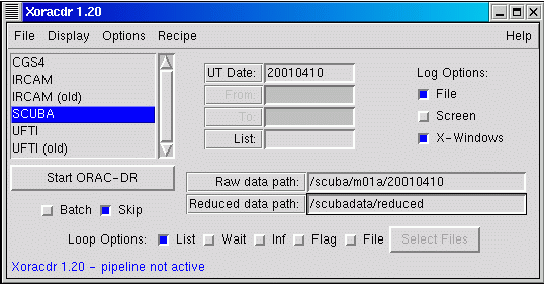
The ORAC-DR pipeline can be controlled from the Xoracdr application. This note describes the user interface of Xoracdr and how to use it.
The ORAC-DR pipeline and Xoracdr software uses a number of environmental variables for configuration. For non-Starlink, non-JAC installation of ORAC-DR, please consult ShellVariables (see appendix) for more information.
To start the Xoracdr application type xoracdr at the prompt:
this will run the Xoracdr setup script and bring up the Xoracdr main window.
Unlike command line control of the ORAC-DR pipeline, configuration of the pipeline options is via the main window. There are therefore only a few command line options
Displays revision information about Xoracdr and the version of Perl it is using.
UT date of observations (defaults to current YYYYMMDD).The UT is required for UKIRT and JCMT instruments as it forms part of the file naming convention for data files.
If you start xoracdr with the ORAC_INSTRUMENT environment variable unset, i.e. you have
not run one of the instrument setup scripts, but have the ORAC_DATA_OUT environment
variable set you may wish to force Xoracdr to honour this using the -honour command
line option.
If your data conforms to the directory naming convention for the instrument, setting up the pipeline is fairly trivial.
Select your instrument from the list box to the left hand side of the application main window and then
enter the UT date of the observations you are reducing into the entry field in the center of
the main window. The raw and reduced data paths should now be set correctly. Enter
the range of frame numbers that you are interested in into the List text entry and push
the Start ORAC-DR button. The pipeline should initalise and start processing you data
immediately.
A more detailed description of the Xoracdr user interface follows for those people who need to do something slightly out of the ordinary, although the interface itself should be fairly self explanatory for those people used to the oracdr command line interface.
The Xoracdr main window is divided into three main sections. At the top is the menu bar with File,
Display, Options, Recipe and Help menus.
When the pipeline is running further data processing can be halted by selecting this menu entry. Control will be returned to the main window, processing can be restarted from the beginning by depressing the Start ORAC-DR button.
When the pipeline is running the processing of further data can be paused by selecting this menu item, a pop-up window will appear with a Resume button. Processing can be re-started by depressing the Resume button in the pop-up.
Selecting this menu option will kill all ORAC-DR related processes, including spawned Starlink monoliths, and will free up any shared memory used by these processes. This will also kill the Xoracdr graphical user interface.
Selecting this checkbox will instruct Xoracdr not to launch the display system. No data will be displayed and GWM (or GAIA) windows will not be launched.
Allows configuration of the ORAC-DR display environment. Selecting this menu item will display a popup window which can be used to edit the current environment and add display directives to the current environment. This popup can also be run as a stand alone application from the command line, e.g.
for more information on how to use the display environment editor see the oracdisp documentation and the documentation on the DisplaySystem.
Allow the pipeline to resume midway through the processing of a group so long as the recipe/instrument supports this behaviour. Default is for the group file to be deleted when a new group is created. When this menu option is selected, the group file is retained. NOTE this option is not currently supported by IRCAM, UFTI and SCUBA recipes.
Do not start algorithm engines. NOTE this will cause the vast majority of recipes to fail.
Do not create an invocation specific temporary directory for the messaging systems
but use whatever directory is the usual default. For ADAM tasks this would mean
that ˜/adam or $ADAM_USER will be used rather than a private ORAC-DR directory.
This should only be used when it is required for ORAC-DR to talk to tasks that were
not started by the pipeline and could lead to possible confusion if multiple pipelines
are running using this flag.
If selected then messages from the Starlink engines will be printed in addition to the normal ORAC-DR messages
Log debug messages to file ORACDR.DEBUG in $ORAC_DATA_OUT.
Turn on perl level warning messages (perl -w). This should be used for debugging
only. If -verbose is also true then full perl diagnostics are turned on (see diagnostics
for more information on this perl pragma).
Is selected the pipeline will make as much noise as possible over errors and pipeline exit.
If selected a window will appear allowing you to enter file paths for the data and calibration root directories, user recipe and primitive directories, the raw and reduced data directories and the instrument calibration directory. If these have already been set, either by selecting an instrument from the listbox or from environment variables you can override the default options using this popup window. NOTE The raw and reduced data paths will be overridden if you select an instrument from the main window listbox after entering values into this window.
This item will remain disabled until an instrument is selected in the main window listbox, after this you are free to select this item from the menu. On selection a window will popup allowing you to enter calibration options for the instrument, see the user guide for your instrument for more information.
Selecting this menu item will ensure that the recipe display window will appear along with the log window. The recipe display window will show the currently executing recipe and a list of the primitives contained in the recipe. The currently running primitive will be highlighted.
This item will be greyed out until an override recipe is selected using the Override
Recipe option further down the Recipe menu. After an override recipe is selected
then this option will become active. If the checkbox is selected then the selected
recipe will override the one specified in the file headers. This override recipe is used
for all data files regardless of header contents or observation mode, so make sure
you only only apply it to appropriate data frames.
This item will remain disabled until an instrument is selected in the main window listbox, after which if selected it will allow you to choose an Override Recipe
Allows you to select and then edit a recipe. The recipe will be saved to
ORAC_RECIPE_DIR, the user recipe directory, which if unset will default to
ORAC_DATA_OUT. Recipes in ORAC_RECIPE_DIR will be run in preference to those in the
instrument recipe directories of the same name.
Selecting an entry from the Help menu will popup a help browser to display the relevant
documentation. At the bottom on the menu there is also an About XORAC-DR entry which will
display the licence terms and conditions for the ORAC-DR software.
At the bottom of the main window is the status bar, this reports the status of the pipeline process. It will display the currently running recipe name, along with other status information.
Between the menu and status bar is the bulk of the main window. On the left is the instrument list box,
this allows you to select the instrument whose data you wish to process. If the ORAC_INSTRUMENT
environment variable is set before starting Xoracdr the instrument will be preselected on
startup.
Once an instrument is selected the user interface will be configured into a default state for the instrument. NOTE it is important to select an instrument before doing any customization of the settings as your changes may be overridden by the default options imposed by setting up an instrument.
At this point the Start ORAC-DR button will become enabled and you can start the pipeline if the configuration options are to your liking.
If not already set using the -ut command line option, and if you are using a loop type where it is
necessary to build the filename using the UT date (i.e. all loop options except file) the UT date of the
observation run can be set via the relevant text entry box in the middle of the main window. NOTE
Doing so will run the instrument setup routine again which will override some environment options
such as ORAC_DATA_IN and ORAC_DATA_OUT.
If ORAC-DR is being run in post-observation mode, the default data detection loop is list. Other loop options can be selected using the series of checkboxes along the bottom of the main window. The available options are
The pipeline will stop once the observations in the list have been reduced.
Waits for data to appear before timing out. Data is reduced and the pipeline waits for the next file.
Do not wait for data. Simply reduce data starting with observation specified by from and
continuing until no more files are present. This is the fastest way of reducing data offline.
Waits for completion files to appear (flags) before processing the data. Data is reduced and the pipeline waits for more data by checking the presence of a flag.
Works much like the list loop option except that looping is carried out over a list of
arbitrarily named files. When the file loop option is selected the Select Files button will be
enabled allowing you to generate this list. NOTE As well as having arbitrary filenames,
files can be added in arbitrary order, this allows you to reduces (for instance) all the
calibration frames for a night first, followed by the actual observations.
There are, in addition, two other options which affect the looping scheme, these being batch and
skip.
Run in batch mode. Precalculate groups before processing data. ‘wait’ loop mode should not be used with this option. NOTE only SCUBA recipes support this option.
Allow the data detection loop to skip missing observations. Default is to stop the loop when an expected file can not be found.
See the DataLoops documentation for more information on looping schemes.
Finally, you can configure the log options from the three checkboxes located to the right hand side of the main window. By default logging is sent to a file and to an Xwindow.
oracdr, oracdisp, oracdr_parse_recipe
Alasdair Allan (aa@astro.ex.ac.uk)
Copyright (C) 1998-2001 Particle Physics and Astronomy Research Council. All Rights Reserved.